Unveiling Bacterial Defense Mechanisms Against Viruses: A Key Insight in the Fight Against Resistance
Antibiotic resistance poses one of the most significant threats to global health in contemporary times. Researchers at Umeå University in Sweden have revealed new insights into this alarming issue, suggesting that the mechanisms bacteria employ to fend off viral attacks may also provide crucial information on how antibiotic resistance develops. With the potential to surpass […]


Antibiotic resistance poses one of the most significant threats to global health in contemporary times. Researchers at Umeå University in Sweden have revealed new insights into this alarming issue, suggesting that the mechanisms bacteria employ to fend off viral attacks may also provide crucial information on how antibiotic resistance develops. With the potential to surpass cancer mortality rates in a few decades, understanding the intricacies of bacterial defense mechanisms is imperative to inform future treatment strategies for antibiotic-resistant infections. The implications of this discovery could herald a new dawn in combating a problem that endangers millions of lives around the world.
The focus of this groundbreaking study lies in the bacterium Staphylococcus aureus, a notorious pathogen known for its potential to cause severe health issues, including septic shock and pneumonia. Alarmingly, certain strains of S. aureus have developed multi-resistance to standard antibiotic treatments, presenting a formidable challenge to public health systems globally. In some regions, the prevalence of multi-resistant S. aureus strains has escalated, with reports indicating that as much as a quarter of infections are due to antibiotic-resistant strains. However, in Sweden, proactive public health measures have managed to keep this figure at about one percent.
Bacteria have coexisted with viruses, specifically bacteriophages, for billions of years, engaging in an evolutionary arms race in which phages strive to infect and kill bacteria while the latter develop sophisticated strategies to resist viral incursions. The phenomenon showcases a delicate balance of power, where the threat of bacteriophages drives bacteria to evolve defenses—an evolutionary strategy that now raises questions concerning their relationship to antibiotic resistance.
The pivotal findings of the Umeå University researchers demonstrate that specific genes within the mobilome of S. aureus confer immunity against phage infections. The mobilome refers to genetic elements that are transferable between bacterial strains, potentially enriching otherwise harmless bacteria with toxic and antibiotic-resistant characteristics. This horizontal gene transfer can lead to the emergence of highly virulent bacterial strains, complicating treatment protocols and further exacerbating global health issues.
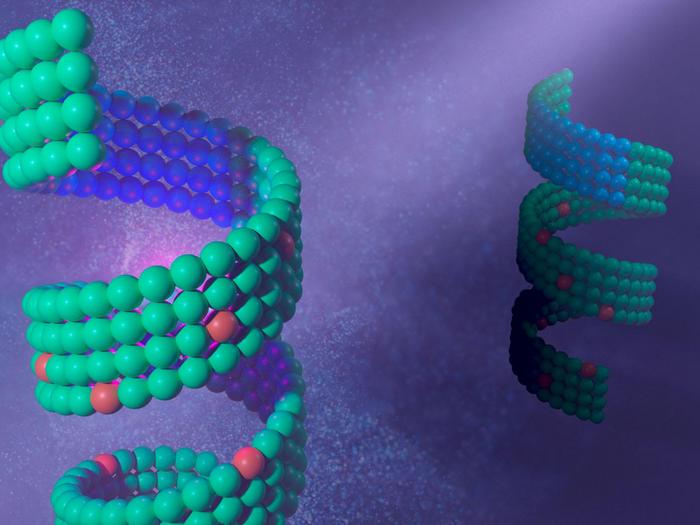
Conventional education about antibiotic resistance has focused primarily on the bacterial mechanisms that resist antibiotics. However, the Swedish research team has unearthed a fascinating new dimension by demonstrating that certain genes in S. aureus mobilome can obstruct the ability of phages to replicate within these bacterial cells. Utilizing state-of-the-art imaging technology, specifically a cryoelectron microscope, they were able to visualize how the bacterial defense proteins interact with components from the phage genome.
These interactions occur at a molecular level, where a key protein—expressed by a gene associated with the mobilome—sufficiently erects a barricade around an essential protein from the phage genome. This blockade critically hampers the phage’s capacity to duplicate its DNA. Consequently, the phage is rendered incapable of launching subsequent attacks on other bacteria, a remarkable find that expands our understanding of bacterial defenses.
This discovery triggers potent ramifications for antibiotic treatment protocols and strategies. By comprehending how resistant bacteria defend themselves against viral agents, researchers can formulate innovative methods to disrupt that defense and thereby reinvigorate the effectiveness of traditional antibiotics. As Mir-Sanchis aptly puts it, the novelty of this mechanism may be a vital piece in the intricate puzzle of antibiotic resistance.
The current research not only unlocks new avenues for combating antibiotic resistance but also sheds light on the evolutionary pressures shaping the bacterial genome. Understanding this dual-functionality—where genes confer resistance to both viral infection and antimicrobial agents—presents significant opportunities for therapeutic advancements. The implications of these findings may lead to pioneering approaches that leverage bacteriophages or other technologies to target resistant bacteria directly.
Moreover, it emphasizes the necessity for continued vigilance and research in the field. Antibiotic stewardship and infection control strategies must evolve in tandem with the understanding of these molecular mechanisms. As bacteria grow increasingly adept at dodging antibiotic effects, an arsenal of strategies—including phage therapy—may provide essential bolstering to our conventional treatment frameworks.
In an age where infections are becoming increasingly resilient to established treatments, the urgency to act has never been clearer. Integrating insights from viral-bacterial interactions offers the potential to reimagine how we categorize and treat bacterial infections, urging a shift towards an increased understanding of microbial ecology.
The foundation laid by this study serves to reinforce the need for interdisciplinary approaches to tackling antibiotic resistance, marrying fundamental biological research with public health initiatives. Comprehending the balance of power between bacteria and the viruses that target them provides crucial knowledge that can be utilized for therapeutic innovations.
As this research opens the door to new scientific inquiries, public health officials and the global medical community are urged to reflect on the findings. The implications emphasize the interconnectedness of ecosystems, highlighting that to adequately address antibiotic resistance, we must also appreciate the roles of viral pathogens in microbial communities.
Moving forward, The Umeå University team’s findings could catalyze new dialogues in research and public health policy, reinforcing a holistic approach to addressing antibiotic resistance. With antibiotic-resistant bacteria posing an existential risk, advancing our understanding of their vulnerabilities and interactions is essential for safeguarding public health now and in the future.
Subject of Research: Cells
Article Title: Phage parasites targeting phage homologous recombinases provide antiviral immunity
News Publication Date: 22-Feb-2025
Web References: http://dx.doi.org/10.1038/s41467-025-57156-3
References: Nature Communications
Image Credits: Umeå University
Keywords: Antibiotic resistance, Staphylococcus aureus, phage therapy, genetic transfer, mobilome, bacteriophages, public health, infection control, viral interactions, molecular mechanisms.
Tags: antibiotic resistance insightsbacterial defense mechanismscombating antibiotic resistancefuture treatment strategiesglobal health challengesimplications of bacterial researchmulti-resistant S. aureus strainspublic health measures in SwedenStaphylococcus aureus health threatsunderstanding antibiotic-resistant infectionsviral attacks on bacteria
What's Your Reaction?